Does my cohort picked the correct number patients? Am I calculating an intersection in the right way? Is that the expected value for treatment duration? It just takes one incorrect parameter to get incoherent results in a pharmacoepidemiological study, and it is very challenging to test calculations on huge and complex databases.
That is why TestGenerator is useful to push a small sample of patients to unit test a study on the OMOP-CDM. It includes tools to create a blank CDM with a complete vocabulary and check if the code is doing what we expect in very specific cases.
This package is based on the unit testing written for the Eramus MC Ranitidine Study.
To install TestGenerator:
# CRAN version
install.packages("TestGenerator")The user can provide an Excel file (link to sample) or a set of CSV files that represent tables of the OMOP-CDM, with a micro population of just 8 patients for testing purposes.
readPatients() will read either Excel or CSVs, and then
saves the data in a JSON file. This is useful if the user wants to
create more than one Unit Test Definitions. If the parameter
outputPath is NULL The files are saved in the
testthat/testCases folder of the package. Alterna
TestGenerator::readPatients(filePath = "~/pathto/testPatients.xlsx",
testName = "test",
outputPath = NULL,
cdmVersion = "5.4")Alternatively, the user can use the functions
readPatients.xl or readPatients.csv
directly.
TestGenerator::readPatients.xl(filePath = "~/pathto/testPatients.xlsx",
testName = "test",
outputPath = NULL,
cdmVersion = "5.4")
TestGenerator::readPatients.csv(filePath = "~/pathto/csv/files",
testName = "test",
outputPath = NULL,
cdmVersion = "5.4",
reduceLargeIds = FALSE)patientCDM() pushes one of those Unit Test Definitions
into a blank CDM reference with a complete version of the vocabulary. If
the pathJSON parameter is NULL,
TestGenerator will look for the JSON test files in the
testthat/testCases folder.
cdm <- TestGenerator::patientsCDM(pathJson = NULL,
testName = "test",
cdmVersion = "5.4")Now the user has a CDM reference with a complete vocabulary and just 8 patients.
filePath <- system.file("extdata/icu_sample_population.xlsx",
package = "TestGenerator")
outputPath <- file.path(tempdir(), "test")
dir.create(outputPath)
TestGenerator::readPatients(filePath = filePath,
testName = "test",
outputPath = outputPath,
cdmVersion = "5.4")
#> ✔ All tables are valid
#> ✔ Unit Test Definition Created Successfully: 'test'
cdm <- TestGenerator::patientsCDM(pathJson = outputPath,
testName = "test",
cdmVersion = "5.4")
#> ! cdm name not specified and could not be inferred from the cdm source table
#> ✔ Standard table(s) in test data: person, observation_period, visit_occurrence, visit_detail, drug_exposure, condition_occurrence and procedure_occurrence
#> ✔ Patients pushed to blank CDM successfully
cdm[["person"]] %>% glimpse()
#> Rows: ??
#> Columns: 18
#> Database: DuckDB 1.5.2 [root@Darwin 24.6.0:R 4.6.0//private/var/folders/wm/s6fjrtt53ld72z03p47nkdvr0000gn/T/RtmpbDvxsa/file41b54c219eb5.duckdb]
#> $ person_id <int> 1, 2, 3, 4, 5, 6, 7, 8
#> $ gender_concept_id <int> 8532, 8507, 8532, 8507, 8532, 8507, 8532, …
#> $ year_of_birth <int> 1980, 1990, 2000, 1980, 1990, 2000, 1980, …
#> $ month_of_birth <int> NA, NA, NA, NA, NA, NA, NA, NA
#> $ day_of_birth <int> NA, NA, NA, NA, NA, NA, NA, NA
#> $ birth_datetime <dttm> NA, NA, NA, NA, NA, NA, NA, NA
#> $ race_concept_id <int> 0, 0, 0, 0, 0, 0, 0, 0
#> $ ethnicity_concept_id <int> 0, 0, 0, 0, 0, 0, 0, 0
#> $ location_id <int> 0, 0, 0, 0, 0, 0, 0, 0
#> $ provider_id <int> 0, 0, 0, 0, 0, 0, 0, 0
#> $ care_site_id <int> 0, 0, 0, 0, 0, 0, 0, 0
#> $ person_source_value <chr> "0", "0", "0", "0", "0", "0", "0", "0"
#> $ gender_source_value <chr> "M", "F", "M", "F", "M", "F", "M", "F"
#> $ gender_source_concept_id <int> NA, NA, NA, NA, NA, NA, NA, NA
#> $ race_source_value <chr> NA, NA, NA, NA, NA, NA, NA, NA
#> $ race_source_concept_id <int> NA, NA, NA, NA, NA, NA, NA, NA
#> $ ethnicity_source_value <chr> NA, NA, NA, NA, NA, NA, NA, NA
#> $ ethnicity_source_concept_id <int> NA, NA, NA, NA, NA, NA, NA, NAThe reference can be used to create a cohort and create unit tests.
test_cohorts <- system.file("extdata",
"test_cohorts",
package = "TestGenerator")
cohort_set <- CDMConnector::readCohortSet(test_cohorts)
cdm <- CDMConnector::generateCohortSet(cdm,
cohort_set,
name = "test_cohorts")
#> ℹ Generating 3 cohorts
#> ℹ Generating cohort (1/3) - diazepam✔ Generating cohort (1/3) - diazepam [1.3s]
#> ℹ Generating cohort (2/3) - hospitalisation✔ Generating cohort (2/3) - hospitalisation [188ms]
#> ℹ Generating cohort (3/3) - icu_visit✔ Generating cohort (3/3) - icu_visit [77ms]
cohortAttrition <- CDMConnector::attrition(cdm[["test_cohorts"]])
excluded_records <- cohortAttrition %>%
pull(excluded_records) %>%
sum()
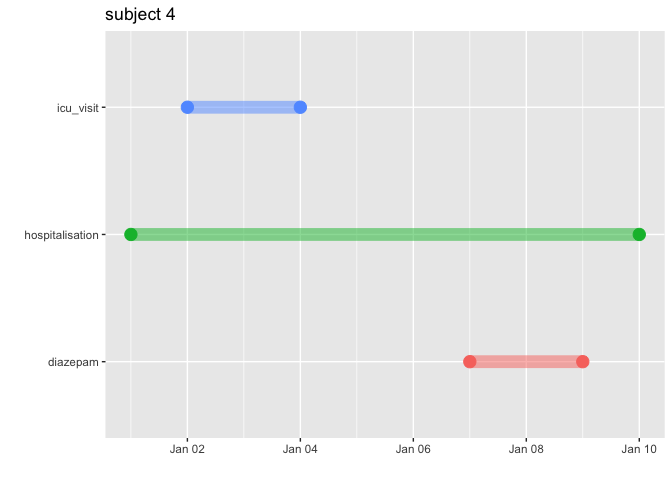
expect_equal(excluded_records, 0)With graphCohort() it is possible to visualise the
timeline for particular patient.
diazepam <- cdm[["test_cohorts"]] %>%
filter(cohort_definition_id == 1) %>%
collect()
hospitalisation <- cdm[["test_cohorts"]] %>%
filter(cohort_definition_id == 2) %>%
collect()
icu_visit <- cdm[["test_cohorts"]] %>%
filter(cohort_definition_id == 3) %>%
collect()
TestGenerator::graphCohort(subject_id = 4, list("diazepam" = diazepam,
"hospitalisation" = hospitalisation,
"icu_visit" = icu_visit))
#> Warning: Using `size` aesthetic for lines was deprecated in ggplot2 3.4.0.
#> ℹ Please use `linewidth` instead.
#> ℹ The deprecated feature was likely used in the TestGenerator package.
#> Please report the issue at
#> <https://github.com/darwin-eu/TestGenerator/issues>.
#> This warning is displayed once per session.
#> Call `lifecycle::last_lifecycle_warnings()` to see where this warning was
#> generated.
#> Warning in geom_segment(aes(x = cohort_start_date, y = cohort, xend =
#> cohort_end_date, : Ignoring unknown aesthetics: fill